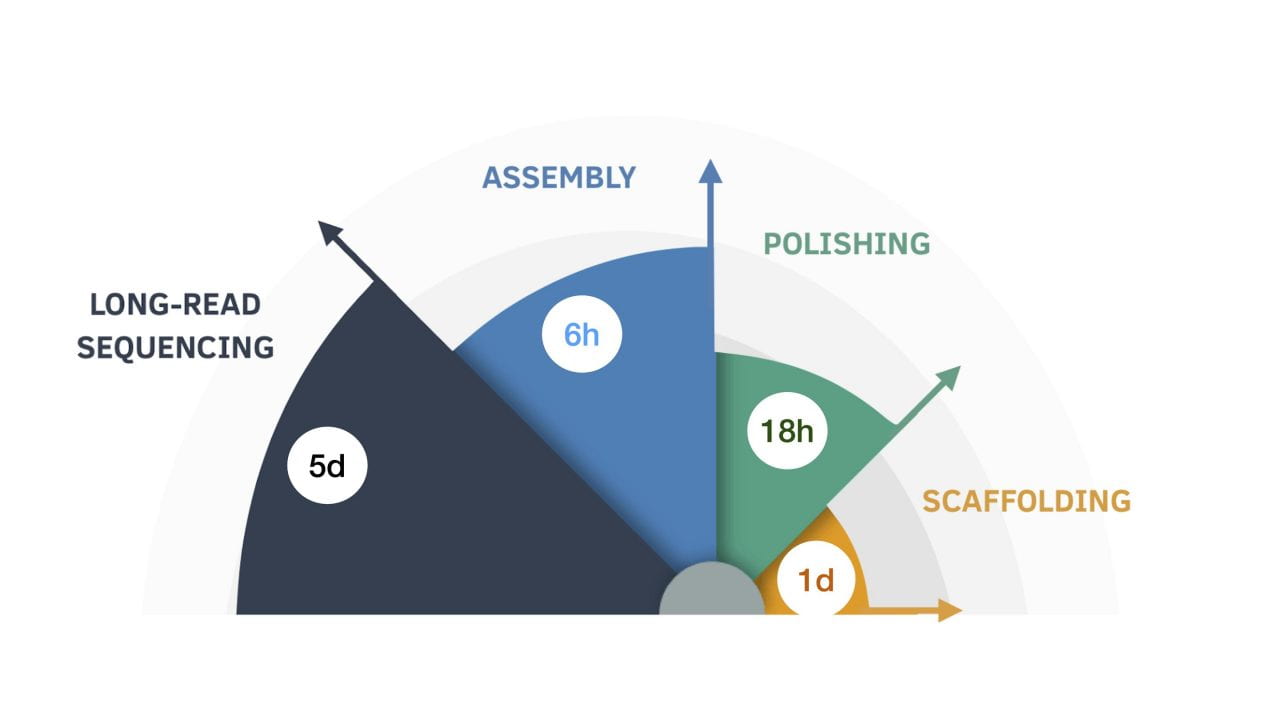
In this diagram, the nine-day genome assembly process is broken down by length of time for each step.
University of California – Santa Cruz | May 4, 2020
It’s only been three years since UC Santa Cruz researchers proved that long-read human genome assembly using the same nanopore technology developed on campus could be done at all. At the time, it was a monumental effort, requiring 150,000 hours of computing time and weeks of work.
About a year later, using the PromethION nanopore sequencer, a similar effort proved significantly faster, cheaper, and easier, clocking in at about a week. “We sequenced eleven human genomes in nine days, which was unprecedented at the time,” said UC Santa Cruz Research Scientist Miten Jain.
Now, researchers at UC Santa Cruz have collaborated on an algorithm designed to accurately and precisely assemble individual, complete human genomes from long-read sequencing data in about six hours and for about $70.
The researchers said they hope their assembler will increase the pace of genomics research and open opportunities. This includes enabling pangenome research to represent the true scale of human diversity, a decidedly more practical pursuit.
Until recently, genomic research has relied exclusively on the reference genome from a single individual selected to represent an entire species. To reflect true human diversity, UC Santa Cruz has embarked on a pangenomic initiative to sequence 350 new, individual human genomes.